Web Tools
Databases
PDIdb (Protein-DNA Interface database) is a repository containing relevant structural information of Protein-DNA complexes solved by X-Ray cristallography and available at the Protein Data Bank (PDB). The database includes a simple functional classification of the protein-DNA complexes that consists of three hierarchical levels (Class, Type and Subtype). This classification has been defined and manually curated by humans based on the information gathered from several sources that included PDB, PubMed, CATH and SCOP. The current version of the database contains only structures with resolution of 2.5 Å or higher. A total of 922 entries are currently available.
Curator: Tomás Norambuena <tanoramb@puc.cl (tanoramb null@null puc NULL.cl)>
Last updated: April, 2010
Web Servers
ANOLEA
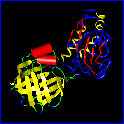 ANOLEA is a server to assess the quality of a three – dimensional protein structure. It uses a statistical potential at the atomic level and gives an energy profile as ouput. Also, the user can choose to have a molecular graphical output representation of the energy profile. The standalone version of the program is also available.
ANOLEA is a server to assess the quality of a three – dimensional protein structure. It uses a statistical potential at the atomic level and gives an energy profile as ouput. Also, the user can choose to have a molecular graphical output representation of the energy profile. The standalone version of the program is also available.
Authors: Melo, F. and Feytmans, E.
dnaMATE
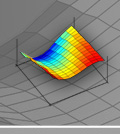 dnaMATE is a server for the calculation of melting temperature for short DNA sequences (15-30 nts) by five different methods. A consensus melting temperature is given, which is based on the results of a comparative study about the similarities and differences of the existing methods that are available for the calculation of melting temperatures. The standalone version of the program is also available.
dnaMATE is a server for the calculation of melting temperature for short DNA sequences (15-30 nts) by five different methods. A consensus melting temperature is given, which is based on the results of a comparative study about the similarities and differences of the existing methods that are available for the calculation of melting temperatures. The standalone version of the program is also available.
Authors: Panjkovich, A., Norambuena, T. and Melo, F.
SAGExplore
![]() SAGExplore is a server for the mapping of SAGE tags against a genomic annotation. Currently, only yeast is available. Xenopus tropicalis and human will be added soon.
SAGExplore is a server for the mapping of SAGE tags against a genomic annotation. Currently, only yeast is available. Xenopus tropicalis and human will be added soon.
Authors: Norambuena, T., Malig, R. and Melo, F.
StAR
 StAR is a web server and standalone program for the statistical comparison of several binary classifiers. This server assesses the statistical significance of the observed differences between two or more classifiers by means of their ROC (Receiver Operating Characteristic) curves. The statistical comparison relies on a non-parametric test and the area under the ROC curves (AUCs). The server is able to perform the simultaneous pairwise comparison of many binary classifiers.
StAR is a web server and standalone program for the statistical comparison of several binary classifiers. This server assesses the statistical significance of the observed differences between two or more classifiers by means of their ROC (Receiver Operating Characteristic) curves. The statistical comparison relies on a non-parametric test and the area under the ROC curves (AUCs). The server is able to perform the simultaneous pairwise comparison of many binary classifiers.
Authors: Vergara, I.A., Norambuena, T., Ferrada, E., Slater A.W, and Melo, F.
WebRASP
 WebRASP is a webserver that uses knowledge-based potentials for scoring RNA structures based on distance-dependent pairwise atomic interactions. This web server allows the users to upload a structure in PDB format, select several options to visualize the structure and calculate the energy profile. The server contains online help, tutorials and links to other related resources. We believe this server will be a useful tool for predicting and assessing the quality of RNA 3D structures.
WebRASP is a webserver that uses knowledge-based potentials for scoring RNA structures based on distance-dependent pairwise atomic interactions. This web server allows the users to upload a structure in PDB format, select several options to visualize the structure and calculate the energy profile. The server contains online help, tutorials and links to other related resources. We believe this server will be a useful tool for predicting and assessing the quality of RNA 3D structures.
Authors: Norambuena, T., Cares, J.F., Capriotti E., and Melo, F.
Outdated
Outdated and unsupported databases and web servers
ProBiSite (http://protein NULL.bio NULL.puc NULL.cl/cardex/databases/pdb/index NULL.html)
ProBiSite (http://protein NULL.bio NULL.puc NULL.cl/cardex/databases/pdb/index NULL.html) (PROtein BInding SITEs) is a database containing all information from the Protein Data Bank (PDB), integrated with the CATH database, and with a ‘home-made’ database that contains useful information about small ligands and protein binding sites in the PDB. This database was built and designed to help in the study of the function/structure relationships in proteins.
Last updated: July, 2002


 (http://www
(http://www